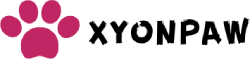AAnyone who thinks cats hate water has clearly never met a fishing cat (Prionailurus viverrinus). These medium-sized (5-17 kg) South and Southeast Asian cats appear equally comfortable in water as on land, capable of swimming long distances both on the surface and underwater. the surface. In captivity, they begin playing in the water from the age of two months, and once adults, about three-quarters of their diet comes from fish.
Unfortunately for these unique cats, the wetlands they depend on are disappearing, and many of those that remain are polluted or claimed by humans, who view the cats as competitors for fish caught and raised in aquaculture. In a study In Thailand, for example, 84% of fishing cats fitted and tracked with radio collars ended up being killed.
Because of a myriad of threats Faced with felines, captive breeding programs have been set up around the world. And this is what allowed scientists to notice at least one threat to animals in captivity: cancer.
Unlike their domestic cousins, captive fishing cats are prone to transitional cell carcinoma (TCC), a cancer of the lower urinary tract. Researchers suspect that their vulnerability to disease may have a genetic basis, as breeding populations are made up of relatively few animals and therefore may have low genetic diversity. But to find disease-causing genes, researchers need genomic tools – tools now available for fishing cats, thanks to a high-quality reference genome published in paper form. bioRxiv pre-publication November 18.
The genome was assembled from PacBio HiFi reads with chromosomal arrangements determined using Hi-C chromatin capture. In total, 96.3 percent of the assembled 2.46 GB genome was assigned to chromosomes, and a BUSCO analysis estimated the sequence to be 93.5 percent complete.
Armed with a robust genome, researchers began the search for candidate TCC-related genes in a cohort of 11 fishing cats, 5 of which were TCC carriers. They first looked for the feline version of eight human genes associated with bladder cancer, observing missense variants in four of these genes in the TCC group:BRCA1, BRCA2, CHEK2And AT M. But BRCA2 stood out, as the team found two missense variants of it in all animals with TBI. However, the variants were also present in half of the control cats, so the team could not conclude that the variants are responsible for the increased risk of TBI in the animals. Additional genomic sampling, which is becoming increasingly affordable, “will help clarify causal risk variants,” the authors conclude.
The study serves as a model for using the vast accumulated knowledge of human diseases to unravel the etiology of diseases in other animals, the authors say, adding that the results of such comparisons could guide mating decisions and warn human caregivers of animals at particularly high risk. future illnesses.
And such discoveries are not one-way street: discoveries about animals can turn on the light on the mysteries of human health. Other researchers have note that feline TCC shares key similarities with human cancer, so further research into TCC in fishing cats may, one day, lead to a more in-depth understanding of this phenomenon cancer in peoplewhich is responsible for approximately 95 percent of all bladder cancers and kills more than 15,000 people in the United States each year.
Finalists:
Northern armyworm (Mythimna separata)
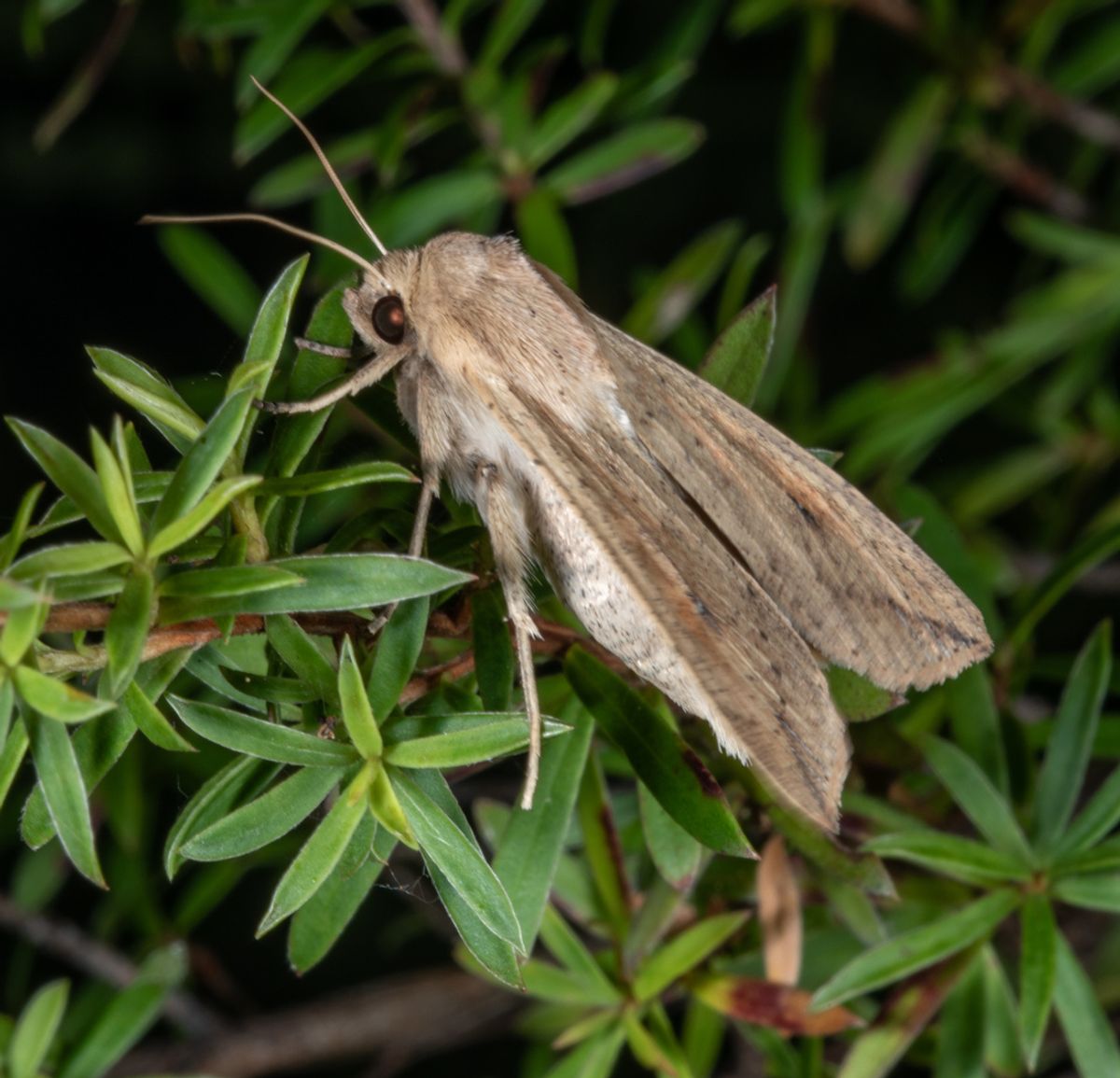

A northern armyworm (Mythimna separata)
Armyworms got a strange nickname because they tend to sprawl in lines and weave their way through grassy habitats – a behavior that can make them a devastating agricultural pest in Asia and Australia. But adults are known for something else: their long nocturnal migrations. Butterflies travel up to 1,400 km in each direction of their journey, orienting themselves without some of the main navigational aids of diurnal migratory species, such as the sun and visual cues. Researchers have long wondered how animals orient themselves and how they fuel such arduous flights. Now, they might finally have the tools needed to unlock these mysteries: A Chinese research team has established a chromosome-level reference genome for the species and designed an efficient gene-editing system that can tinker with this genome, according to a study. published on December 20 in Cell Reports. The 170.3 GB genome, estimated to be 98% complete, was assembled from PacBio long reads, with chromosomes determined via Hi-C. The team then silenced genes suspected of being involved in magnetoreception, which disrupted their ability to orient themselves at night. The team also used their gene-editing system to explore moth coloration; eliminate the gene pale resulted in lighter butterflies, while eliminating ebony leads to darker cuticles. “Successful KO of pale And ebony not only validates their respective roles in the formation of the melanization trait, but also demonstrates the feasibility of a high-efficiency genetic manipulation system for future functional studies and application of genetic management of this destructive pest species,” write the authors in the article.
Costate mountain snail (Oreohelix idahoensis)
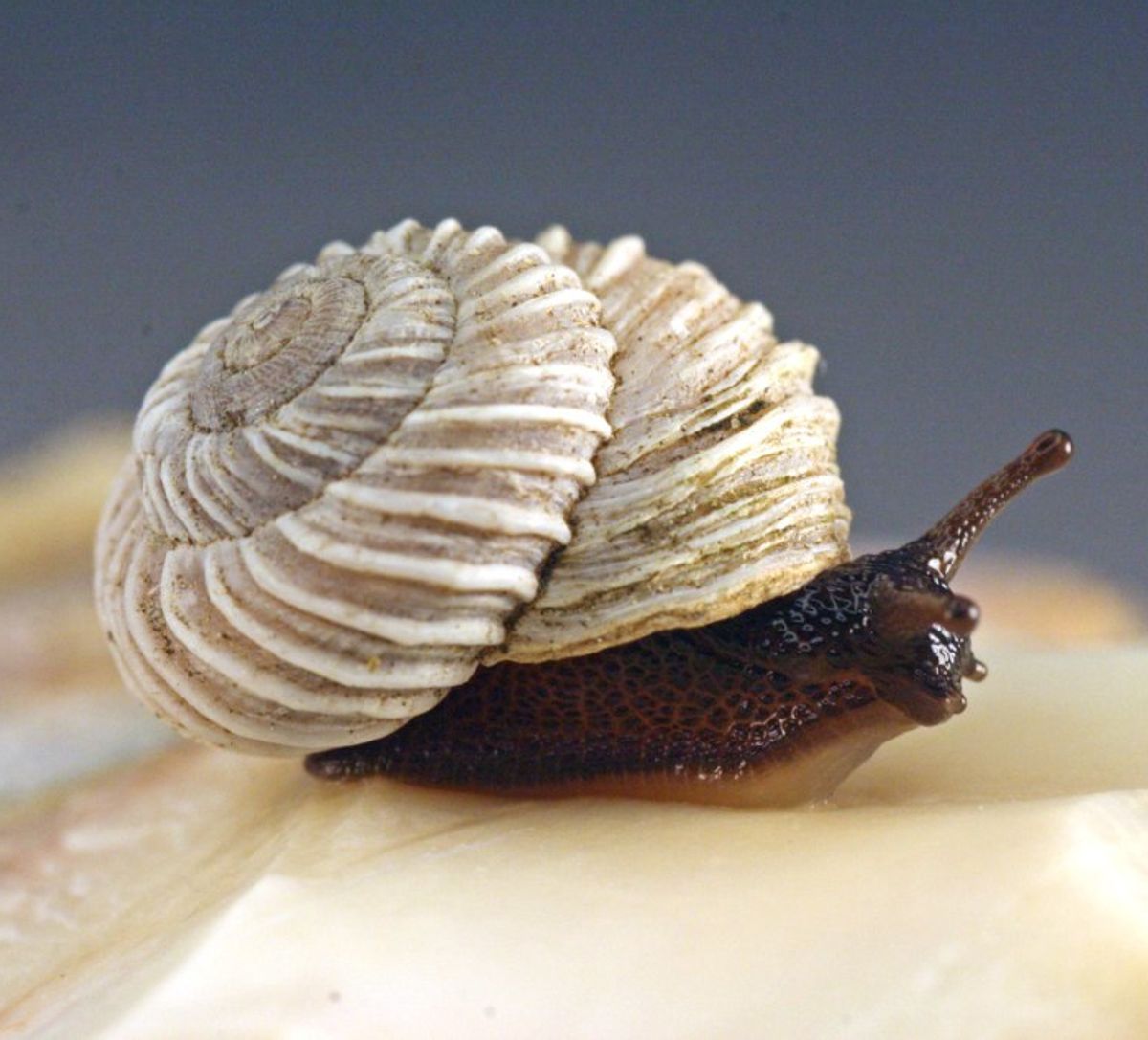

A costate snail (Oreohelix idahoensis)
Limestone rocks like limestone and marble are essential ingredients in cement production. But humans aren’t the only animals that love sucking rocks: limestone habitats are often home to diverse and specialized ecosystems. Mountain snails, in particular, appear to have repeatedly radiated into calcareous habitats, with many species now specialized in calcareous substrates. One such species in the costate mountain snail (Oreohelix idahoensis), which is endemic to calcareous habitats of the northwestern United States. To better understand how this species adapted to its carbonate-rich environment, the researchers assembled the snail’s 5.4 GB genome from PacBio long reads and 10x Genomics linked reads. As they report on December 2 in BMC genomics, it is the largest mollusk genome assembled to date. It is also the most repetitive, with long terminal repetitions accounting for over 57% of sequences. Indeed, repeat content was estimated to be 2 to 3 times higher in the costate mountain snail than in two related non-calcareous snails. This discovery “is unprecedented in mollusk genomics and sheds new light on how transposable element content can vary between molluscs,” the authors write. “The genomic resources reported here will enable further studies of the genomic mechanisms underlying the specialization of limestone rocks and the evolution of transposable element content between molluscs.”